DNA makes a wonderful material for creating structures at the nanoscale. It can be folded and shaped into three dimensions mimicking the traditional Japanese art of paper origami but at scales one million times smaller. Nano DNA shapes made in this way mimic the intricate 3D structures of biological proteins and have a large range of potential applications such as biosensing and metamaterials.
My PhD work focuses on using DNA origami for improving the sensitivity of nanopore technology. Nanopores are crucial to a sensing method for detection of single molecules by passing them through tiny pores (1-100nm in diameter) and measuring the changes in ionic current they cause as they move through the pore. They hold great promise as a high throughput technique for a wide range of applications such as DNA sequencing and single molecule protein detection. A great challenge in the field is the ability to control on nanometre length-scales the three dimensional geometry and surface chemistry of the nanopore. DNA origami allows us to synthesise structures on this length scale with almost arbitrary control of geometry and nanometre positioning of functional chemical groups.

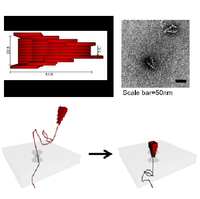
Formation of a DNA origami nanopore. Top image: computer design of the DNA nanopore. Each rod represents a double helix and dimensions are marked in nanometres. Bottom image: electron microscope side-view image of DNA nanopores formed by self-assembly.
We designed a DNA origami nanopore shape as shown in the figure above with its constriction of 7.5nm [1]. To form an architecture for single molecule sensing we docked this DNA origami nanopore with a solid state nanopore approximately 15nm in size (see figure below). This was achieved by applying a voltage across the solid state nanopore so that the DNA origami was electrophoretically trapped into the solid state nanopore. The process is reversible since an opposite voltage expels the DNA origami from the solid state nanopore. We are now working on methods for designing DNA origami nanopore structures that mimic the behaviour of proteins which control the movement of small molecules across cell membranes.

- DNA Origami Nanopores
Nicholas A. W. Bell, Christian. R. Engst, Marc Ablay, Giorgio Divitini, Caterina Ducati, Tim Liedl, and Ulrich F. Keyser
Nano Letters 2012 12 (1), 512-517
Nicholas Bell
NanoDTC PhD Student Cohort 2009